This page contains links to GitHub projects. I still do not readily use GitHub for all of my work, but will post and update 'R packages' as I develop them.
|
ssmc
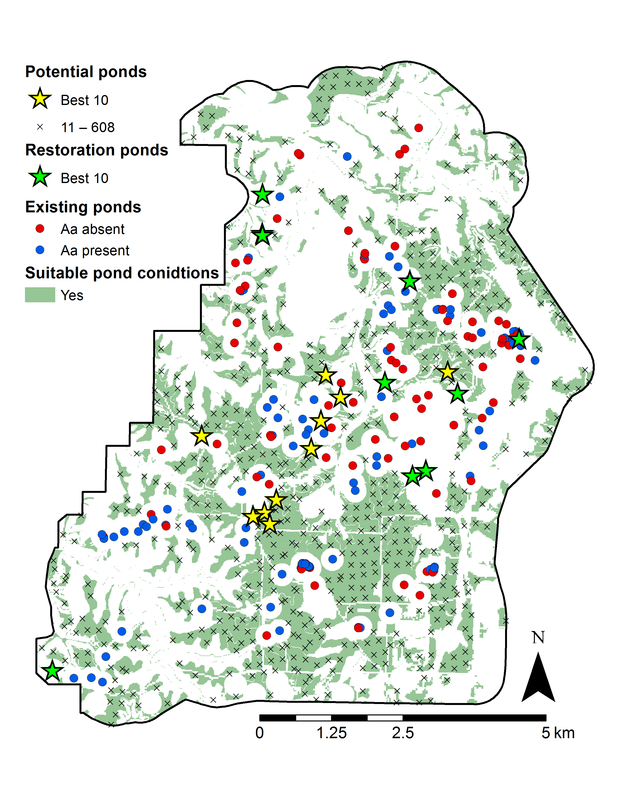
(GitHub Link) 'ssmc' stands for Source-Sink Monte Carlo, and is an R package for assessing source-sink status of populations based on a demographic connectivity model and Monte Carlo sampling. With this package you can assess the the source-sink status of individual populations as well as the importance of populations to metapopulation persistence. Additionally, you can determine the best locations on a landscape to create new habitat or identify existing habitat that if restored, would have the greatest benefit to the metapopulation. These methods were originally developed for the assessment of pond breeding amphibians, but should be adaptable to any species with discrete habitat requirements. Detailed documentation of how to use this package can be found in here. This package is early in development, so please notify me of issues or bugs that arise. Also let me know if there are functions or features that you would like to see added. |
|
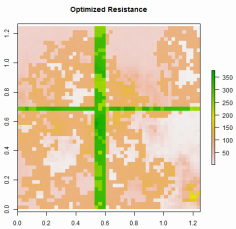
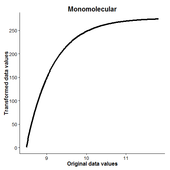
ResistanceGA
(GitHub Link) This package is an improvement on the methods originally described in Peterman et al. (2014) and expanded in Peterman (2018). Optimization of resistance surfaces now utilizes genetic algorithms, which provide a much more flexible and powerful framework for resistance optimization. Both continuous and categorical surfaces can be optimized using these methods. Additionally, multiple resistance surfaces can be optimized simultaneously to produce novel resistance surfaces. This R package allows you to optimize using circuit-based resistance distances (calculated in CIRCUITSCAPE), or cost distances calculated from least cost paths. A vignette/tutorial demonstrating the functions in the package can be found here (NOTE: The Vignette is outdated!). Please contact me if you have issues with these functions, struggle with interpretation, or would like to see other features added. If using CIRCUITSCAPE, it is highly recommended to install Julia and the CIRCUITSCAPE Julia package. General instructions for doing so can be found here. |
|
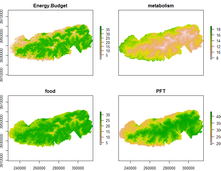
biophys
(GitHub Link) An R package to create spatially-explicit biophysical models. This package is being developed in collaboration with Matt Gifford and should be considered an alpha version. Documentation is currently very limited. Functions are written for specific, personal needs, but the package will become more developed in the future as we expand this model and functions to address more questions. |
|
ResistanceOptimization
(GitHub Link) This package uses the functions and methods originally described in Peterman et al. (2014). Since the publication of the Molecular Ecology paper, I have vastly improved the methods used to optimize resistance surfaces, which are now implemented in ResistanceGA. |
|
iBdry
(GitHub Link) This code and sample data can be used to batch process iButton data, with an emphasis on determining drying and filling dates of wetlands. A manuscript describing the use of iButtons for determining wetland inundation is currently in revision. In the future, this code will be made into a simple R package. |